Cryptic species have for long time been considered as purely a taxonomical challenge. However, in the last decade it has been shown that their recognition has also consequences for several other biological disciplines. Recently, their importance for understanding certain evolutionary processes has been highlighted. Most prominent among these is the process of stasis, which features also very strongly in paleontology. Essentially, stasis means that species did not change for extended periods of time, often millions of years. In the case of cryptic species, this is not only a single species, but several species, which retained the morphology. Last year, we could show that species of the annelid genus Stygocapitella did not change for up to 140 million years. To put this into perspective, at that the time dinosaurs still walked the earth.
Now we published a paper in PeerJ together with colleagues from the US, UK and Portugal using Stygocapitella as a model system to understand genomic causes of morphological stasis. For our study, we used three Atlantic species of Stygocapitella, which have been separated from each other for 18 and 6 million years, respectively. We exploited a population genomic approach based on RADseq resulting in an analysis of more than 250,000 loci of their genomes. This allowed us to address the following hypotheses with respect to stasis at the genomic level; something not possible with paleontological data.
(1) bottlenecks and founder effects reduce genetic variation, thus resulting in morphological similarity
(2) morphological similarity results from recent admixture
(3) shared genetic variation due to incomplete lineage sorting and ancient admixture underlies morphological similarity
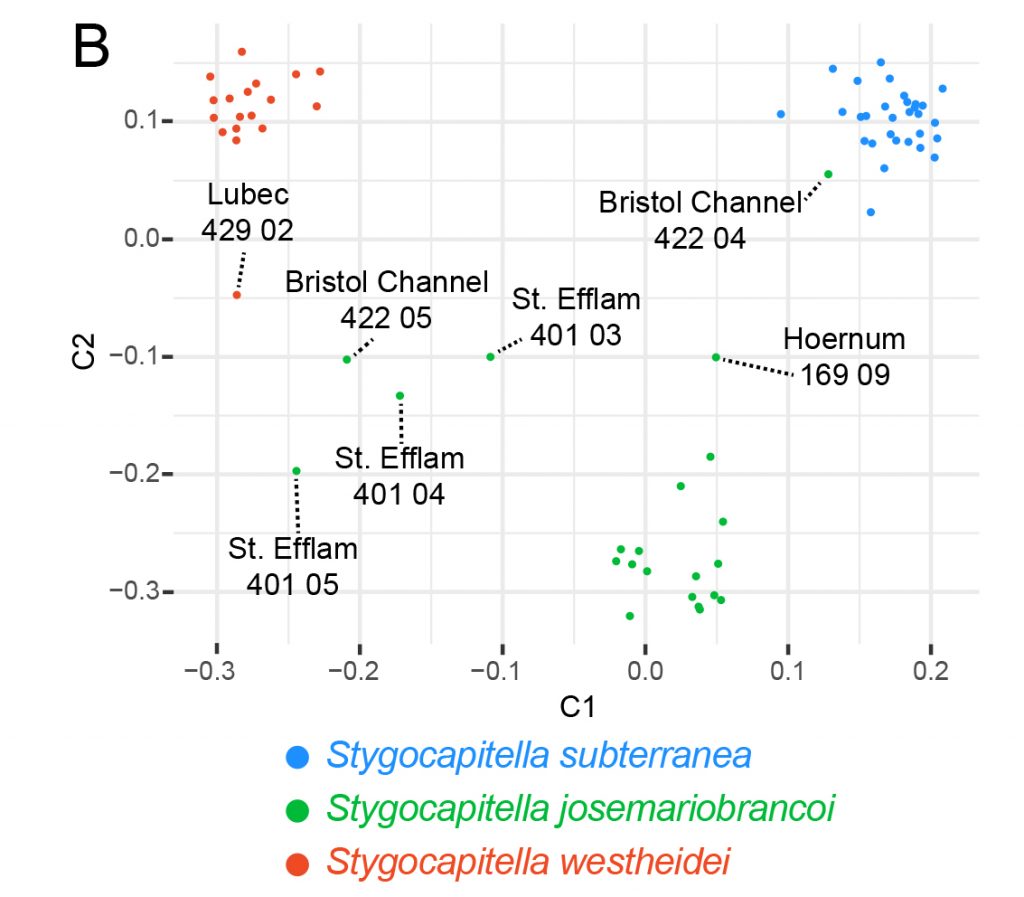
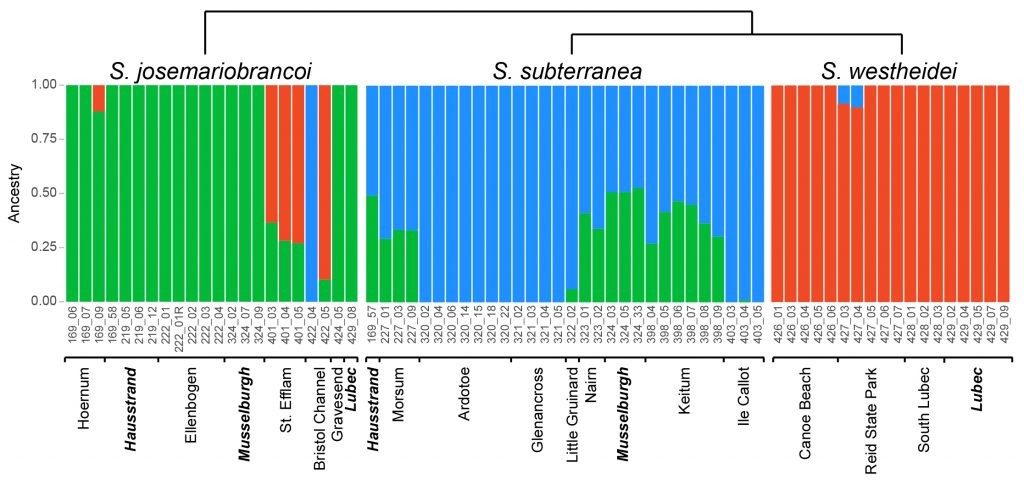
Our results showed that the three species share genomic information. However, further testing revealed this shared genomic information was neither the results of reoccurring bottlenecks and founder effects nor of recent admixture. The shared signal stems from either ancient admixture or more likely incomplete lineage sorting. While we could not directly show that the morphological similarity is directly caused by the standing shared genetic signal, it nonetheless provides additional indication for this hypothesis as a possible cause for morphological. In any case, ongoing recent hybridization mixing parts of the gene pools of the morphological identical species can be excluded. The quest for understanding the causes of no morphological change despite ongoing evolution at other levels of the biological system still needs further investigations and is ongoing, but we could add a little puzzle piece to it.
![]()

2 Comments on “What causes species not to change despite ongoing evolution?”