During his PhD Jose was interested in the evolution of the cryptic species complexes in the annelid genus Stygocapitella, which occurs at sandy beaches around the world. Part of his thesis also comprised population-genomic studies to understand the underlying genomic background of morphological stasis. During the phylogenomic studies using RADseq data he encountered problems with missing loci in the datasets as the species had been separated from each for millions of years.
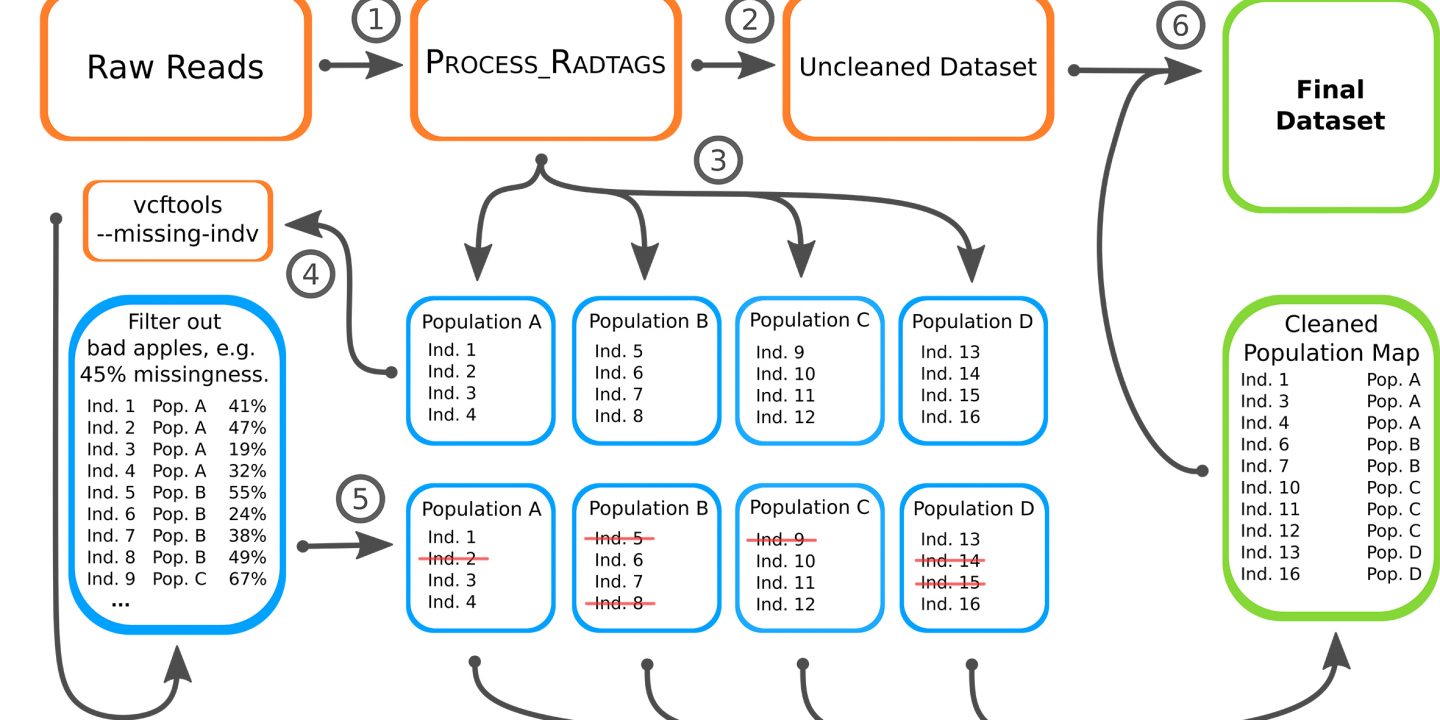
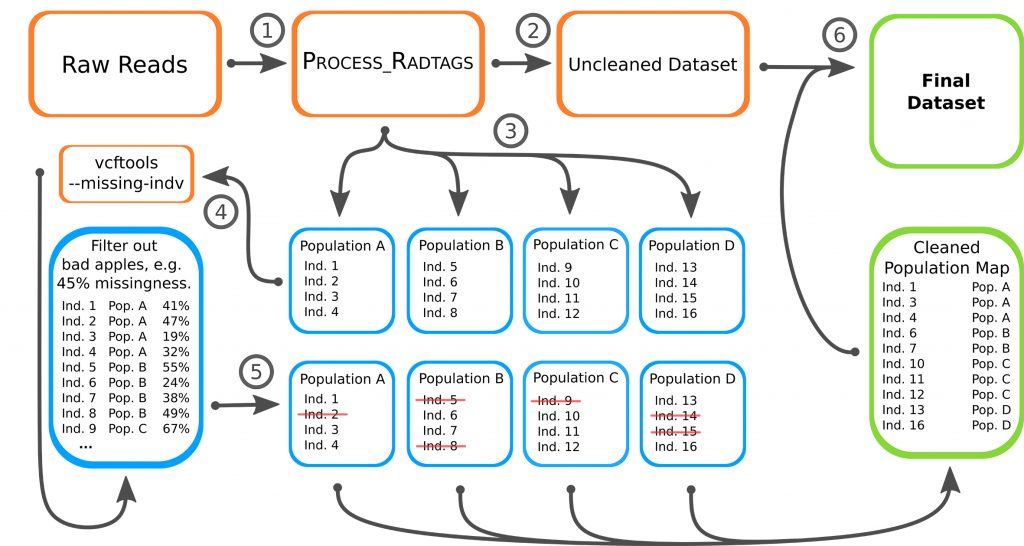
To improve the data he thought about a new approach to reduce missingness in the data. He called it the “bad apple” approach as its principle is based on detecting the species in a population that has the highest degree of missingness and which are above certain threshold. This is analysis is first conducted on the population level only. Then the results from the population level are applied to the entire data of all three species.
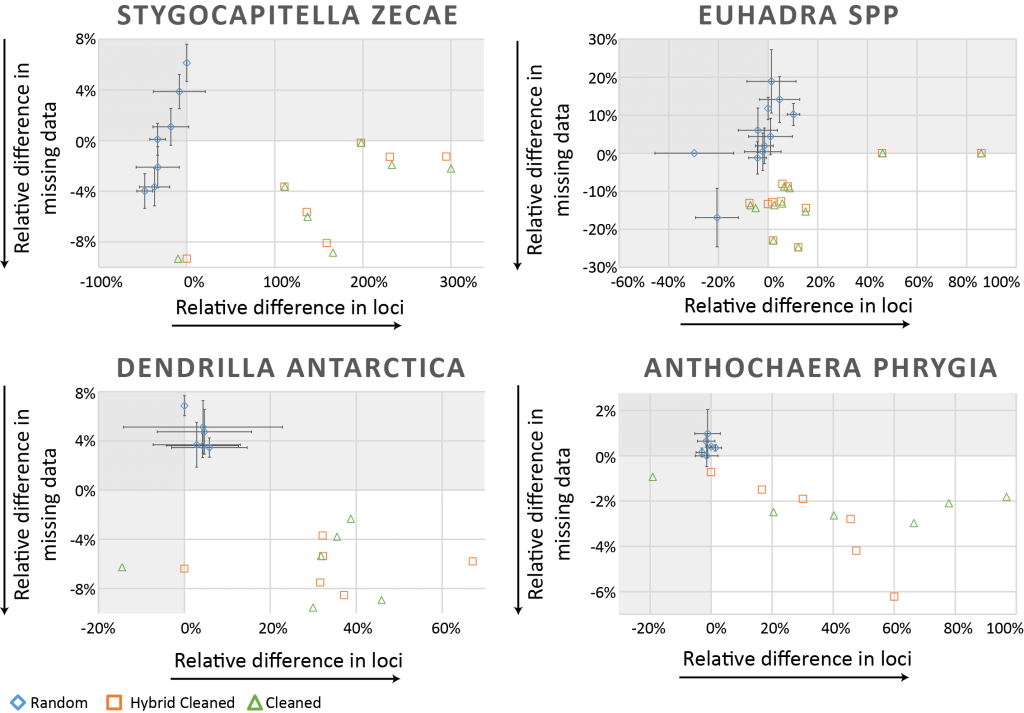
Jose reached out to the community to test the approach on other datasets as well. In the end, besides the worm Stygocapitella zecae a snail genus (Euhadra), a bird species (Anthochaera phrygia) and a sponge species (Dendrilla antarctica) were included in the study. In all four datasets, the approach proved to be very fruitful. At least either the number of loci or the degree of missingness could be improved, but in most cases both could be achieved. Randomly deleting the same number of specimens did not achieve the same effect and often had the opposite effect.
The paper has been published in Methods in Ecology and Evolution and is a cooperation with the lab of Julian Catchen at University of Illinois at Urbana-Champaign. Moreover, many of the analyses were done by Marius Maurstad, whose was a very interested Bachelor student at that time and is doing his Master thesis now. He did it just out of curiosity for the subject.
![]()