Today, I would like to present to the paper by Tianzhu Xiong and James Mallet about “On the impermanence of species: The collapse of genetic incompatibilities in hybridizing populations” in Evolution. The topic of the paper is not my area of research, which are deep level phylogeny, evolutionary genomics, cryptic species and stasis. Accordingly, the paper was quite a mouthful for me. However, it was worth working my way through the paper. Nonetheless, I would like to highlight that if anything in this blog today is not presented correctly, it is due to my limitations and not due to the authors.
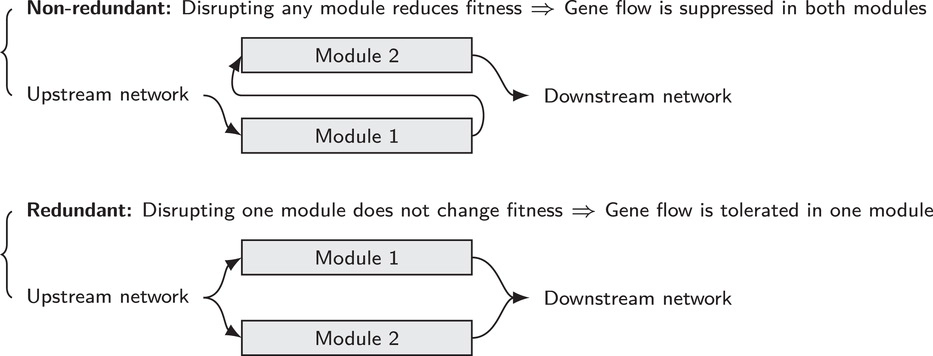
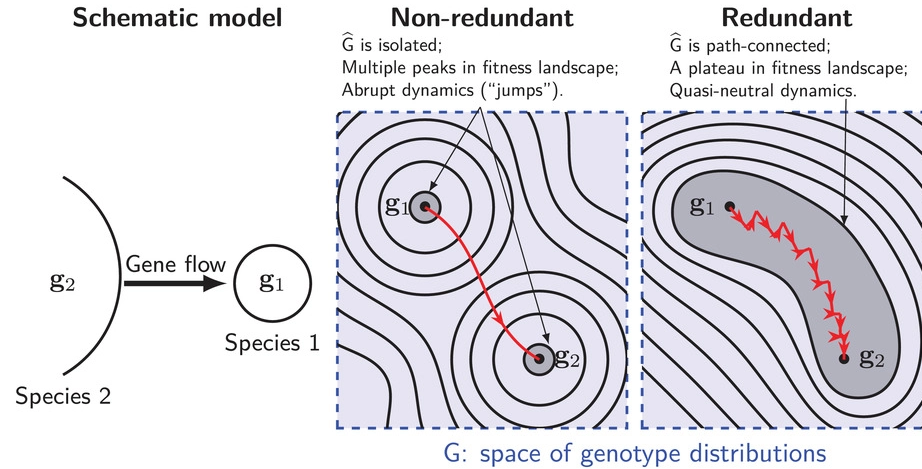
In this paper, the authors address the topic how incompatibilities, which have build up in species pairs after divergence, can collapse in hybrids and essentially leading to a hybridization of the species in at least some populations. This phenomenon is also known as the incompatibility collapse. The authors look at a very specific case. They “argue that redundancy of genetic pathways can strongly affect the dynamics of intrinsic incompatibilities”. From this, they conclude that “higher redundancy decreases the stability
of incompatibilities during hybridization, but also increases tolerance of incompatibility polymorphism within species”. To test these hypotheses, the authors use simulation studies, where they assess the effect of different scenarios such as different levels of redundancy and recombination.
The authors have several interesting findings in their paper such as that the level of redundancy show very different behaviors in relation to genetic drift. In non-redundant systems, the mean time to an eventual collapse is shorter at strong drift, while it vice versa in redundant systems. Generally, in redundant systems the time to collapse is faster than in non-redundant systems. However, for me the most interesting results was that independent of degree of redundancy and other parameters, in redundant systems the incompatible alleles of a loci remained in the populations to at least a small degree even though the alleles are incompatible with each other (see figure below). This is possible due to the redundancy in the system.
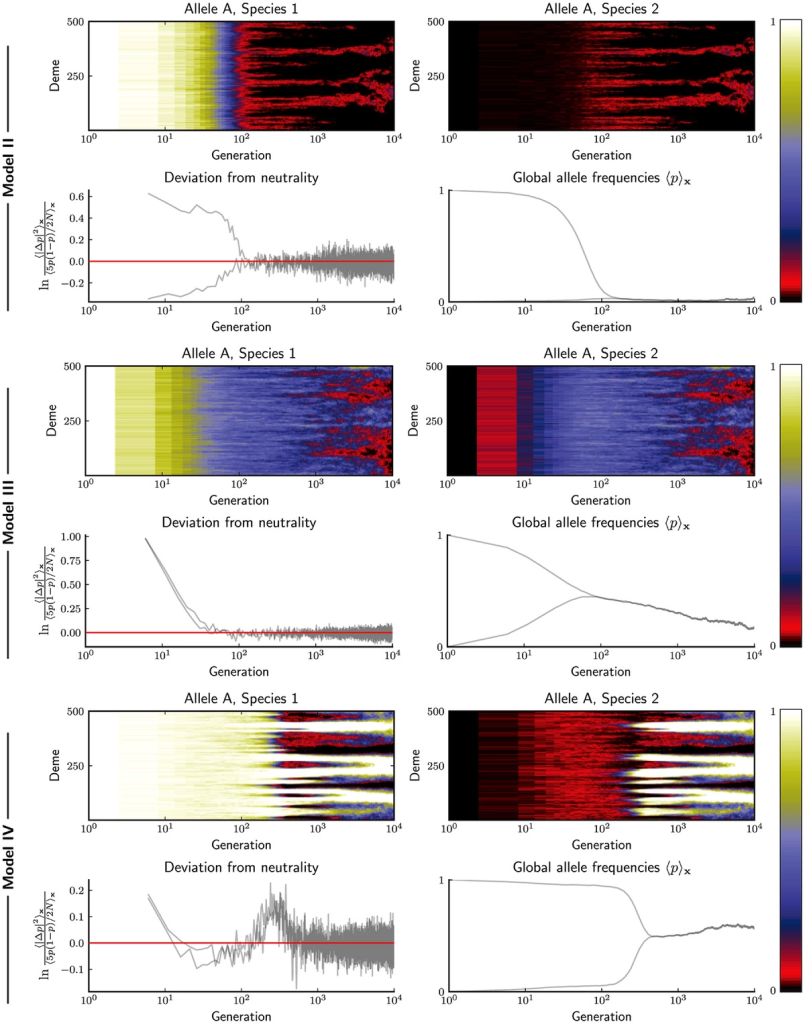
species and the higher the level of redundancy (increasing from Model II to IV) the more polymorphism remains, but even at the lowest levels of redundancy (Model II) a small degree of polymorphism remains at a stable low level.
Why is this now interesting with respect to stasis? Several hypotheses explaining morphological stasis assume that in some populations we can actually observe adaptation to the local environmental differences, but these are either driven back to the original phenotype or that they are swamped by it when an ephemeral population get in contact again with the major population (e.g., source-sink relationships, fluctuation around a mean). If redundancy in systems allows for an increased level of polymorphism and more importantly even keeping incompatible alleles in the system, it is much easier to understand how the original phenotype be stable even though in certain populations local adaptation takes place. If the species with stasis have higher level of redundancy then original alleles could remain in the populations even if they are incompatible with the new alleles and a pushback to the old state is much easier as it can work on the standing genetic variation. The nice aspect of this is that the hypothesis can be easily tested. A prediction would be that species or traits with stasis have a higher degree of duplicated/redundant genes and/or of pseudogenes in their genomes than non-static species. Hence, as said above getting these inspiring ideas was by itself worth reading a paper outside of my comfort zone and shows the value of it. I would like to thank the authors for this inspiring paper.
![]()