On the 10th Day of Advent, we open the door on our calendar to a paper that brought me more than a little bit of joy to read earlier this year, Yoshida et al’s “Time-series transcriptomic screening of factors contributing to the cross-tolerance to UV radiation and anhydrobiosis in tardigrades”. Tardigrades are an unendingly hilarious series of questions upon questions, and it feels a little like, when you’ve just gotten a grasp on how things work within the phylum, they slide through your hands once again. This paper contributes greatly to my continued gleeful, engaged bafflement with tardigrades which I’m sure will continue for years to come, so it felt like a perfect time to share it.
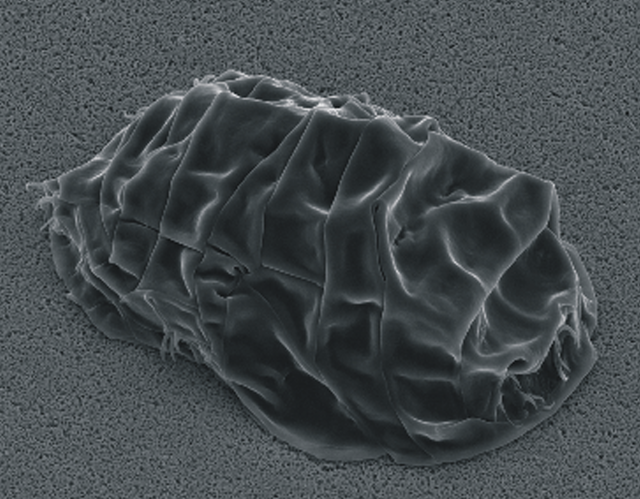
Tardigrades can tolerate extreme environments through a process called cryptobiosis, where they reduce themselves to an inactive, suspended state. More than just hibernation or torpor, which are the resting states we see in many animals, cryptobiosis is a “little death”, a much more complete reduction in life function. In tardigrades, this manifests specifically as “anhydrobiosis” – cryptobiosis by desiccation. It allows them to survive extreme temperatures, high pressure and even exposure to the vacuum of space, which has not only made them celebrities in the pop science eye in recent years, but also a subject of some fascination amongst biologists.
Trying to work out the molecular mechanism behind anhydrobiosis has been a puzzle that scientists have been working on for a long time across animals, plants, fungi and bacteria. Tardigrades, in fact, are not the only animal to engage in anhydrobiosis! However, they are the only ones we know of where a sugar called trehalose does not seem to be particularly important. That suggests that the mechanism of tardigrade anhydrobiosis is unique even within the animals. A number of tardigrade-specific protein families have previously been identified: but these aren’t found across all tardigrades either, suggesting that we’re missing some elusive smoking gun, a protein family that helps us understand tardigrade anhydrobiosis across the whole group.
Yoshida et al. started by using transcriptomes to study which genes were expressed before and after one species of tardigrades were exposed to UVC radiation. As transcriptomes are “snapshots” of which genes are expressed at a point in time, if we change one variable and compare transcriptomes, we can work out what genes are involved in what processes. They found a new gene, called g12777, which seemed to be expressed a lot more in response to the stress, and followed a similar expression pattern to some genes we know are also involved in anhydrobiosis. What’s more, it was also found across most of the other species of tardigrade genomes they have access to, which suggests it could have been found in the ancestor of all tardigrades. It was expressed in the Golgi apparatus, an important part of many animal cells that “packages” the proteins to be sent off elsewhere in the cell, which also suggests that this part of the cell might be more important in surviving desiccation than we had previously thought.
So could this g12777 be the smoking gun we’ve been looking for?
Well… maybe! We’ll need to look at how it is expressed, and where, and why, in the other tardigrades we can find it in. We’ll need to repeat these transcriptome expression studies in lots of other species, to see if it is doing the same job, and consider other kinds of functional gene studies. It might be that it performs different roles in other tardigrades, or isn’t expressed as strongly in them. But it feels like we might be one step closer to understanding anhydrobiosis in tardigrades, and that’s like completing a line in a very challenging Rubik’s Cube on Christmas morning. An achievement worth celebrating.
![]()