The year 2021 was full of unexpected happenings and opportunities for both students, as in me Marianna Khodabandehlou and my fellow student, Sine Hagestad. We both were doing our last semester in our Biosciences degree and wanted to gain some hands-on experience before we began our Masters. We were very happy to be chosen from all the other 100 candidates when we applied to the UiO:Life Science, for the summer project at the Natural History Museum (NHM) which involved assessing the biodiversity of marine invertebrates in the Oslo fjord. We were honoured to work together with professor Torsten Hugo Struck and technical assistant Ana Teresa Capucho from June 2021 to September 2021 on the Artsdatabanken Project connected to our summer project.
The summer project involved a lot of fieldwork and many hours in the laboratory, where we learned to combine all our knowledge from our bachelor’s to real-life techniques and data analysis. Torsten taught us patiently about the fieldwork techniques for capturing the samples (Figure 1) and how important cooperating together was to make good and correct observations.

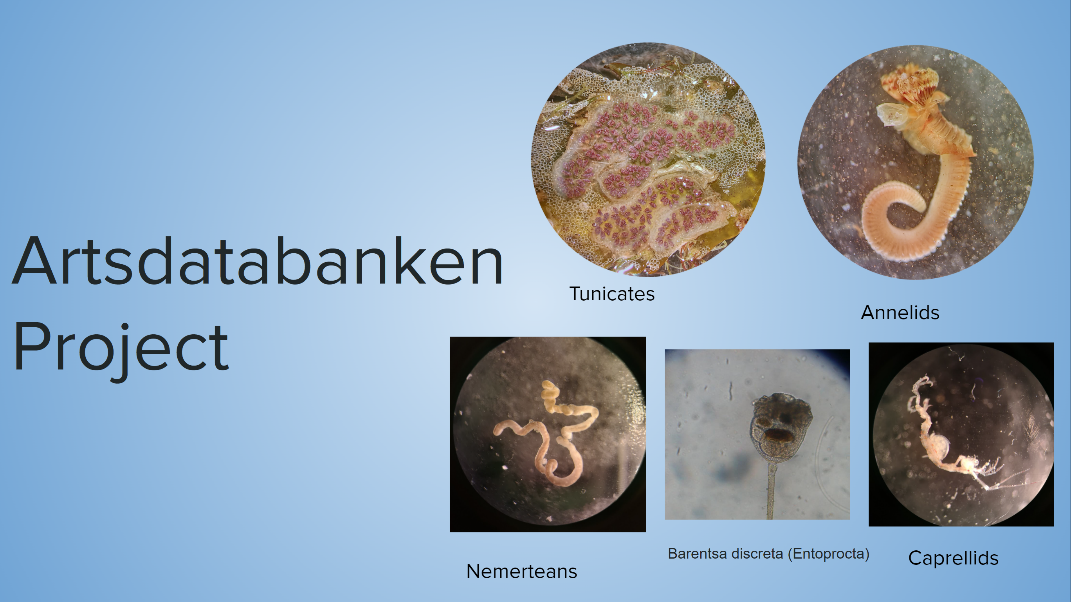
In addition, we weren’t afraid of getting our hands dirty and doing our best to locate and recognize the target species for the project. Figure 2 shows the microscopy pictures captured by me and Sine in the laboratory in order to record the invertebrates we had captured before we prepared them for DNA Barcoding in the DNA Laboratories at NHM. DNA barcoding is commonly used to learn more about the taxonomy of unknown organisms such as the species we have captured in our fieldwork (Wilson et al., 2018). The preparation involved learning to work with separating the samples in ethanol and formalin, as well as all the safety measures and technical details that we had to know before working further with DNA Barcoding.

Working in the DNA Lab with Sine and Ana Teresa, our technical assistant and helpful guide in this project, made us realize how small missteps can cause a whole day of experiments to be thrown off balance and how to work precisely and carefully in all the steps to reach a good result in our barcoding. Sequencing a short fragment of the mitochondrial cytochrome c oxidase subunit I (COI) gene helped us compare the DNA fragments from the captured invertebrates with an online library of available DNA barcodes of known taxonomy (Wilson et al., 2018) and to further evaluate with highest accuracy which one was the observed species.
After we had completed our work at the Natural History Museum with Ana Teresa and Torsten, me and Sine prepared a scientific poster for the UiO:Life Science Summer Projects Final Meeting, together with a presentation for the guests (Figure 3).
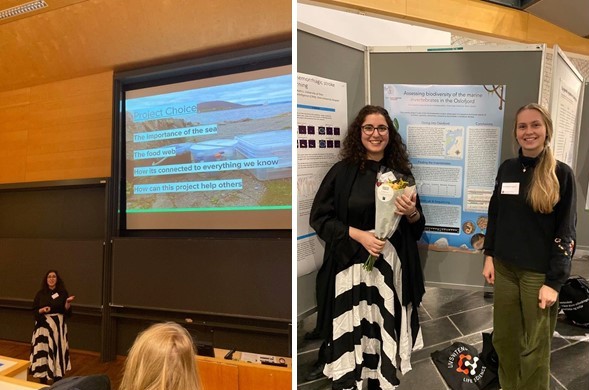
We were more than honoured to win an award for the popular science poster at this event and receive good feedback from our work (Figure 4).

We have been very lucky to be involved in such an interesting project, with Torsten Struck and Ana Teresa Capucho, who gave us not only a wonderful learning experience but also inspired us for our future master’s choices. I will be continuing with a master in Toxicology and Environmental Sciences working with arctic copepods and Sine will be continuing with a master in Marine Biology and Limnology. We will miss this project and this team, but we are excited to see what the future holds for us.
Marianna Khodabandehlou
Read more: Life Sciences: Closing event for the summer students
References:
– Wilson, J. J., Sing, K-W., & Jaturas, N. (2018). DNA Barcoding: Bioinformatics Workflows for Beginners. In B. D. Roitberg (Ed.), Reference Module in Life Sciences Elsevier. https://doi.org/10.1016/B978-0-12-809633-8.20468-8
![]()
